If you use IPython you can usually find out where functions are defined with by appending ?? to the function. For example:
>>> from sklearn.svm import SVC
>>> svc = SVC()
>>> svc.score??
Signature: svc.score(X, y, sample_weight=None)
Source:
def score(self, X, y, sample_weight=None):
"""Returns the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy
which is a harsh metric since you require for each sample that
each label set be correctly predicted.
Parameters
----------
X : array-like, shape = (n_samples, n_features)
Test samples.
y : array-like, shape = (n_samples) or (n_samples, n_outputs)
True labels for X.
sample_weight : array-like, shape = [n_samples], optional
Sample weights.
Returns
-------
score : float
Mean accuracy of self.predict(X) wrt. y.
"""
from .metrics import accuracy_score
return accuracy_score(y, self.predict(X), sample_weight=sample_weight)
File: ~/miniconda/lib/python3.6/site-packages/sklearn/base.py
Type: method
In this case it's coming from the ClassifierMixin so this code can be used with all classifiers.
https://github.com/scikit-learn/scikit-learn/blob/master/sklearn/base.py#L310
https://ipython.readthedocs.io/en/stable/interactive/python-ipython-diff.html#accessing-help
Answer from Jacques Kvam on Stack OverflowI am looking for SVM code without using any library. (Raw SVM code for Sentiment analysis). Can anyone help me in this?
NEED HELP WITH SVM KERNEL CODE IN PYTHON FROM SCRATCH
machine learning - Implementation of SVM for classification without library in c++ - Stack Overflow
python - Support vector machine from scratch - Stack Overflow
Videos
Hi everyone
Sorry if what I am asking is elementary but I am new to machine learning and I have been asked to build a SVM classifier with Gaussian or Polynomial Kernel which solves the Dual Quadratic Problem from scratch without importing it directly from any library like sci-kit learn.
I understand the math behind it but since I am new to coding I am unable to find a comprehensive YT video since everyone just imports it from sci-kit learn, I tried finding books but they were from a long time ago and didn't have python implementation, I tried GitHub code, but in two instances that I found it, it was all over the place.
Can anyone link me up to a simple code or let me know where to search, I will be extremely appreciative of you.
I will join to most people's advice and say that you should really consider using a library. SVM algorithm is tricky enough to add the noise if something is not working because of a bug in your implementation. Not even talking about how hard is to make an scalable implementation in both memory size and time.
That said and if you want to explore this just as a learning experience, then SMO is probably your best bet. Here are some resources you could use:
The Simplified SMO Algorithm - Stanford material PDF
Fast Training of Support Vector Machines - PDF
The implementation of Support Vector Machines using the sequential minimal optimization algorithm - PDF
Probably the most practical explanation that I have found is the one on the chapter 6 of the book Machine Learning in action by Peter Harrington. The code itself is on Python but you should be able to port it to C++. I don't think it is the best implementation but it might be good enough to have an idea of what is going on.
The code is freely available:
https://github.com/pbharrin/machinelearninginaction/tree/master/Ch06
Unfortunately there is not sample for that chapter but a lot of local libraries tend to have this book available.
Most cases SVM is trained with SMO algorithm -- a variation of coordinate descent that especially suits the Lagrangian of the problem. It is a bit complicated, but if a simplified version will be ok for your purposes, I can provide a Python implementation. Probably, You will be able to translate it to C++
class SVM:
def __init__(self, kernel='linear', C=10000.0, max_iter=100000, degree=3, gamma=1):
self.kernel = {'poly' : lambda x,y: np.dot(x, y.T)**degree,
'rbf' : lambda x,y: np.exp(-gamma*np.sum((y - x[:,np.newaxis])**2, axis=-1)),
'linear': lambda x,y: np.dot(x, y.T)}[kernel]
self.C = C
self.max_iter = max_iter
def restrict_to_square(self, t, v0, u):
t = (np.clip(v0 + t*u, 0, self.C) - v0)[1]/u[1]
return (np.clip(v0 + t*u, 0, self.C) - v0)[0]/u[0]
def fit(self, X, y):
self.X = X.copy()
self.y = y * 2 - 1
self.lambdas = np.zeros_like(self.y, dtype=float)
self.K = self.kernel(self.X, self.X) * self.y[:,np.newaxis] * self.y
for _ in range(self.max_iter):
for idxM in range(len(self.lambdas)):
idxL = np.random.randint(0, len(self.lambdas))
Q = self.K[[[idxM, idxM], [idxL, idxL]], [[idxM, idxL], [idxM, idxL]]]
v0 = self.lambdas[[idxM, idxL]]
k0 = 1 - np.sum(self.lambdas * self.K[[idxM, idxL]], axis=1)
u = np.array([-self.y[idxL], self.y[idxM]])
t_max = np.dot(k0, u) / (np.dot(np.dot(Q, u), u) + 1E-15)
self.lambdas[[idxM, idxL]] = v0 + u * self.restrict_to_square(t_max, v0, u)
idx, = np.nonzero(self.lambdas > 1E-15)
self.b = np.sum((1.0 - np.sum(self.K[idx] * self.lambdas, axis=1)) * self.y[idx]) / len(idx)
def decision_function(self, X):
return np.sum(self.kernel(X, self.X) * self.y * self.lambdas, axis=1) + self.b
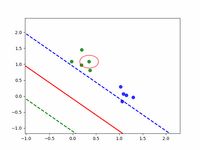
In simple cases it works not much worth than sklearn.svm.SVC, comparison shown below

For more elaborate explanation with formulas you may want to refer to this ResearchGate preprint. Code for generating images can be found on GitHub.
It seems like the implementation of your code is correct. At such small margins, you should not worry about minor increases in your cost.
The increase in cost occurs when your learning rate multiplied by your gradient "overshoots" the optimal value. In this example, it happened by an extremely small amount so I would not worry about it.
If you are curious about why the cost increase at all, we first have to ask why shouldn't it? Gradient descent points in the direction which minimizes our loss. However, if our learning rate is large enough, we can shoot past the optimal value and end up with a larger cost! This is what your code essentially did, except at an extremely small and negligible scale.
I figured out what was wrong. When calculating the hinge loss, I should have used X @ w -b instead of X @ w + b; this affects how the bias term being updated during gradient descent. From my experience with these code, SGD reforms better than batchGD in general, and requires less hyper-parameter tuning. But theoretically, if a learning rate with decay is used for batchGD, it should perform no worse than SGD.
Hi everyone
Sorry if what I am asking is elementary but I am new to machine learning and I have been asked to build a SVM classifier with Gaussian or Polynomial Kernel which solves the Dual Quadratic Problem from scratch without importing it directly from any library like sci-kit learn.
I understand the math behind it but since I am new to coding I am unable to find a comprehensive YT video since everyone just imports it from sci-kit learn, I tried finding books but they were from a long time ago and didn't have python implementation, I tried GitHub code, but in two instances that I found it, it was all over the place.
Can anyone link me up to a simple code or let me know where to search, I will be extremely appreciative of you.
You did not search thoroughly enough or you don't know how to search
The most valuable skill that you urgently need to develop is how to search for information and be less dependent on handouts
whenever you have a question like this, you fire up the search engine of your choice and you type:
"implement [YOUR ARCHITECTURE HERE] from scratch"